Academics
| Degree | University/Institution |
|---|---|
| Ph.D. | School of Biosciences and Bioengineering, IIT Bombay. |
| MSc. (Organic Chemistry) | Ravenshaw College, Cuttack |
Work Experience
| Position | University/Organisation | Period |
|---|---|---|
| Scientist-F | Inistitute of Life Sciences, Bhubaneswar | 2024 - Present |
| Scientist-E | Inistitute of Life Sciences, Bhubaneswar | 2019 – 2023 |
| Scientist-D | Inistitute of Life Sciences, Bhubaneswar | 2015 – 2018 |
| Ramalingaswami Fellow Scientist | Inistitute of Life Sciences, Bhubaneswar | 2011-2016 |
| Post-Doctoral Fellow | University of Texas Health Science Center at Houston | 2006-2011 |
Awards & Recognisation
| Details |
|---|
|
Research
| Details |
|---|
Laboratory of Bacterial Cell Division and Antimicrobial Resistance
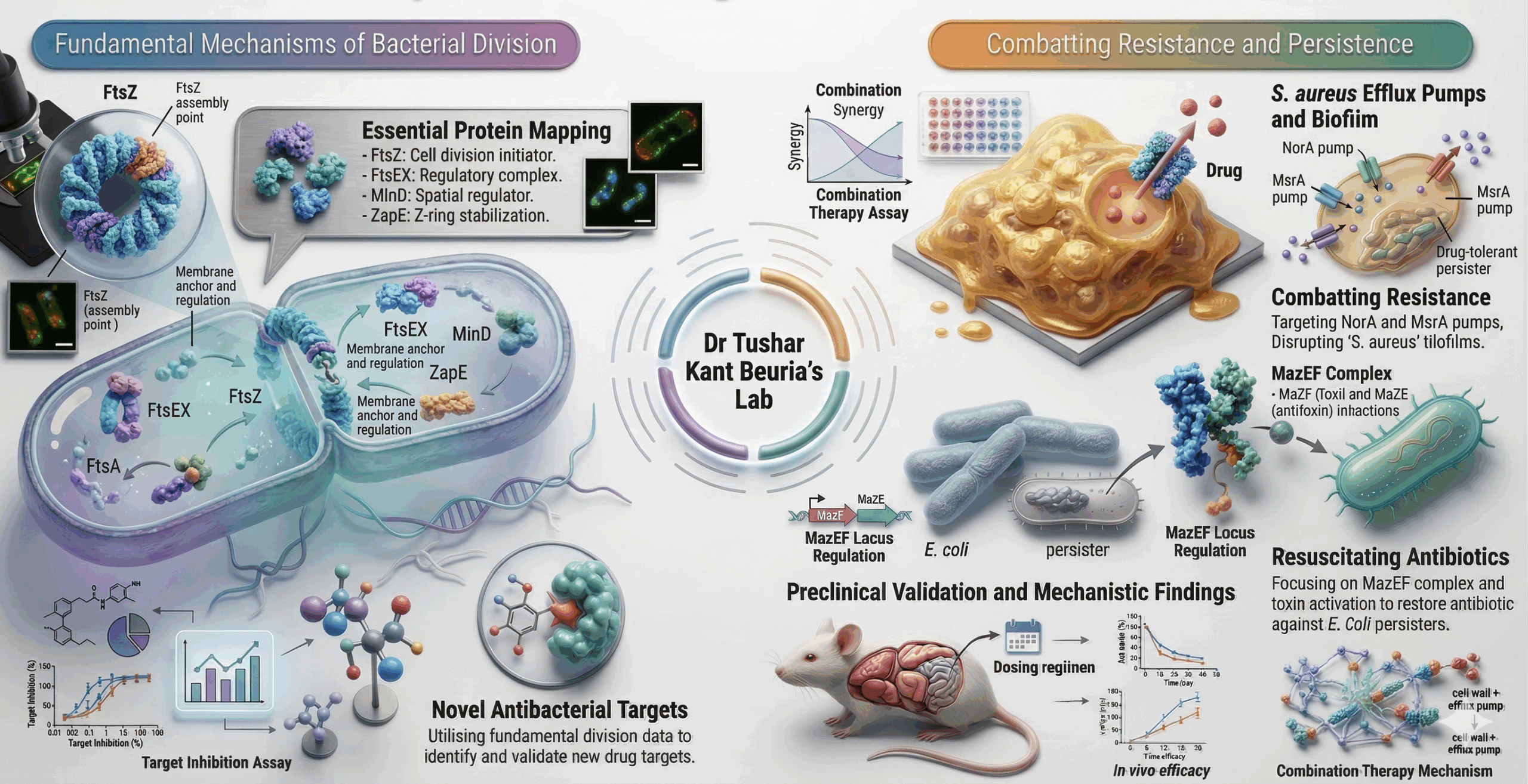
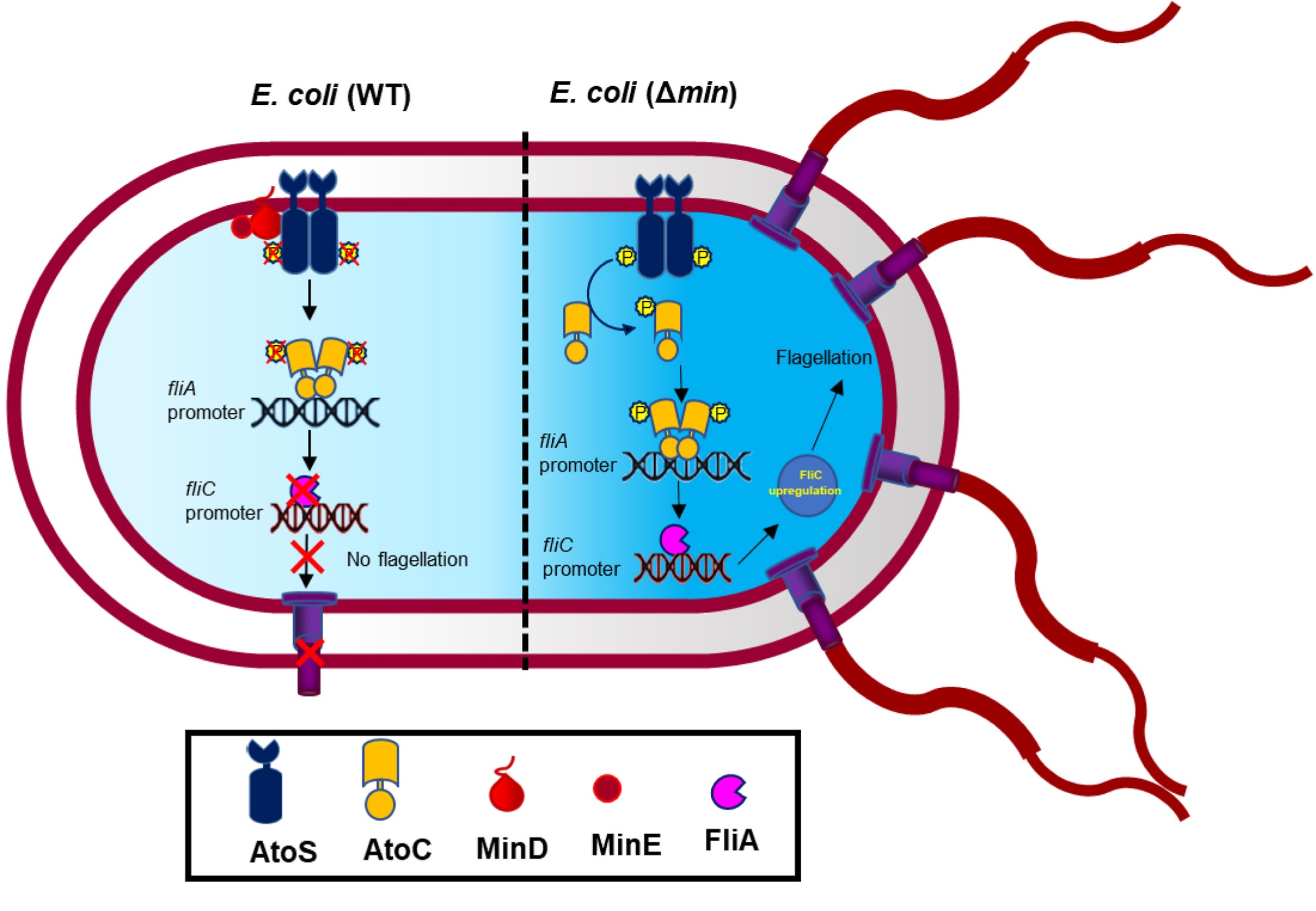
Research: 1. Bacterial Cell Division MachineryUnderstanding the molecular mechanisms of bacterial cell division has been a central focus of my laboratory’s research programme. Bacterial cell division is an essential and highly conserved process orchestrated by the divisome, a supramolecular machine anchored by FtsZ. This prokaryotic tubulin homolog polymerises into a dynamic Z-ring at the midcell. Given that FtsZ and its associated proteins are highly conserved across most bacteria yet absent in humans, the divisome represents a compelling and largely underexploited target for novel antibacterial drug discovery. Over the years, my group has mapped critical protein-protein interaction interfaces within the divisome, characterised the FtsZ interdomain cleft as a druggable vulnerability, defined FtsEX-dependent coordination of septal peptidoglycan synthesis, and identified novel small-molecule binding sites on FtsZ, including unexpected interactions with Doxorubicin (Biochem J, 2015) and aztreonam’s targeting of ZapE (FEBS Lett, 2023). Most recently, we demonstrated that MinDE independently governs bacterial flagellation beyond its canonical division role (J Biol Chem, 2024), expanding our understanding of Min protein biology. Our current focus is on exploiting these mechanistic insights to develop potent, selective inhibitors of divisome assembly as next-generation antibacterial agents. |
Publications
| Details |
|---|
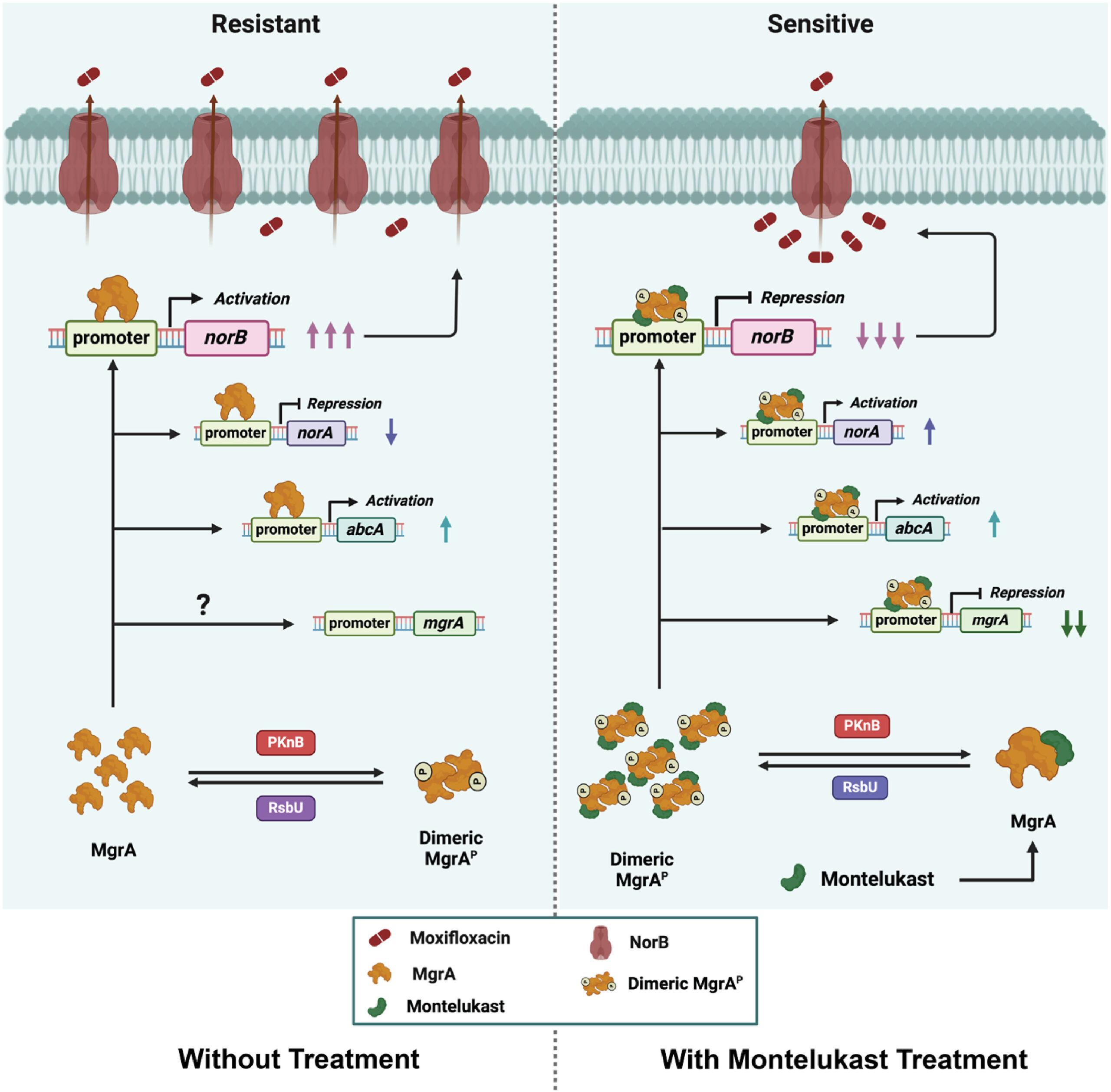
2026Efflux-Mediated Resistance in Staphylococcus aureus: Challenges and Opportunities for Therapeutic Intervention Ojha S, Beuria TK, Indian Journal of Microbiology, 1–30 (2026), DOI: 10.1007/s12088-026-01545-9 2025Efflux pump modulation by Montelukast and its roles in restoring antibiotic susceptibility in multidrug-resistant Staphylococcus aureus Ojha S, Sinsinwar S, Chatterjee P, Biswal S, Pradhan P, Beuria TK, EBioMedicine, 114, 105658 (2025) PubMed: 40157128
10058-F4 Mediated inhibition of the biofilm formation in multidrug-resistant Staphylococcus aureus Dodia H, Ojha S, Chatterjee P, Beuria TK, Biofilm, 8, 100307 (2025), DOI: 10.1016/j.bioflm.2025.100307
Inhibition of efflux pumps by FDA-approved drugs oxiconazole and sertaconazole restores antibiotic susceptibility in multidrug-resistant S. aureus Ojha S, Chatterjee P, Beuria TK, Antimicrobial Agents and Chemotherapy, 69(9), e00320-25 (2025), PubMed: 40757818 2024The bacterial division protein MinDE has an independent function in flagellation Pradhan P, Taviti AC, Beuria TK, Journal of Biological Chemistry, 300(4), 107117 (2024), PubMed: 38403244
2023The cell division protein ZapE is targeted by the antibiotic aztreonam to induce cell filamentation in Escherichia coli Guru A, Taviti AC, Sethy M, Ray S, Dixit A, Beuria TK, FEBS Letters, 597(23), 2931–2945 (2023), PubMed: 37857499
FtsE, the nucleotide binding domain of the ABC transporter homolog FtsEX, regulates septal PG synthesis in E. coli Mallik S, Dodia H, Ghosh A, Srinivasan R, Good L, Raghav SK, Beuria TK, Microbiology Spectrum, 11(3), e02863-22 (2023), PubMed: 37014250
2022Long-read 16S-seq reveals nasopharynx microbial dysbiosis and enrichment of Mycobacterium and Mycoplasma in COVID-19 patients Prasad P, Mahapatra S, Mishra R, Murmu KC, et al. Molecular Omics, 18(6), 490–505 (2022), DOI: 10.1039/d2mo00044j
Isolation and characterization of five SARS-CoV-2 strains of different clades and lineages circulating in Eastern India Singh B, Avula K, Chatterjee S, Datey A, et al. Frontiers in Microbiology, 13, 856913 (2022, DOI: 10.3389/fmicb.2022.856913
MBZM-N-IBT, a novel small molecule, restricts Chikungunya virus infection by targeting nsP2 protease activity in vitro, in vivo, and ex vivo De S, Ghosh S, Keshry SS, Mahish C, Mohapatra C, Guru A, et al. Antimicrobial Agents and Chemotherapy, 66(7), e00463-22 (2022), DOI: 10.1128/aac.00463-22
Kanamycin-Mediated Conformational Dynamics of E. coli outer membrane protein TolC Pattanayak BS, Dehury B, Priyadarshinee M, Jha S, Beuria TK, Integrative Structural Biology of Proteins and Macromolecular Assemblies (Book Chapter) (2022), Search on Google Scholar
2021Targeting the achilles heel of FtsZ: the interdomain cleft Pradhan P, Margolin W, Beuria TK, Frontiers in Microbiology, 12, 732796 (2021), DOI: 10.3389/fmicb.2021.732796
Identification of multipotent drugs for COVID-19 therapeutics with the evaluation of their SARS-CoV-2 inhibitory activity Kumar S, Singh B, Kumari P, Kumar P, et al. Computational and Structural Biotechnology Journal, 19, 1998–2017 (2021), DOI: 10.1016/j.csbj.2021.04.014
Clinical, virological, immunological, and genomic characterization of asymptomatic and symptomatic cases with SARS-CoV-2 infection in India Chatterjee S, Datey A, Sengupta S, Ghosh A, et al. Frontiers in Cellular and Infection Microbiology, 11, 725035 (2021), DOI: 10.3389/fcimb.2021.725035
Biochemical characterization of an E. coli cell division factor FtsE shows ATPase cycles similar to the NBDs of ABC-transporters Mallick S, Kumar A, Dodia H, Alexander C, Vasudevan D, Beuria TK, Bioscience Reports, 41(1), BSR20203034 (2021), DOI: 10.1042/BSR20203034
Kanamycin-mediated conformational dynamics of Escherichia coli outer membrane protein Tol Pattanayak BS, Dehury B, Priyadarshinee M, et al. Frontiers in Molecular Biosciences, 8, 636286 (2021) DOI: 10.3389/fmolb.2021.636286
Comparative profile of ocular surface microbiome in vernal keratoconjunctivitis patients and healthy subjects Vishwakarma P, Mitra S, Beuria T, Barik MR, et al. Graefe’s Archive for Clinical and Experimental Ophthalmology, 259(7), 1925–1933 (2021), DOI: 10.1007/s00417-021-05109-z
Nanopore 16S rRNA sequencing reveals alterations in nasopharyngeal microbiome and enrichment of Mycobacterium and Mycoplasma in patients with COVID-19 Mahapatra S, Mishra R, Prasad P, Murmu KC, et al. medRxiv (Preprint) (2021) DOI: 10.1101/2021.11.10.21266147
Isolation and Characterization of SARS-CoV-2 strains circulating in Eastern India Singh B, Avula K, Chatterjee S, Datey A, et al. bioRxiv (Preprint) (2021), DOI: 10.1101/2021.12.13.472526
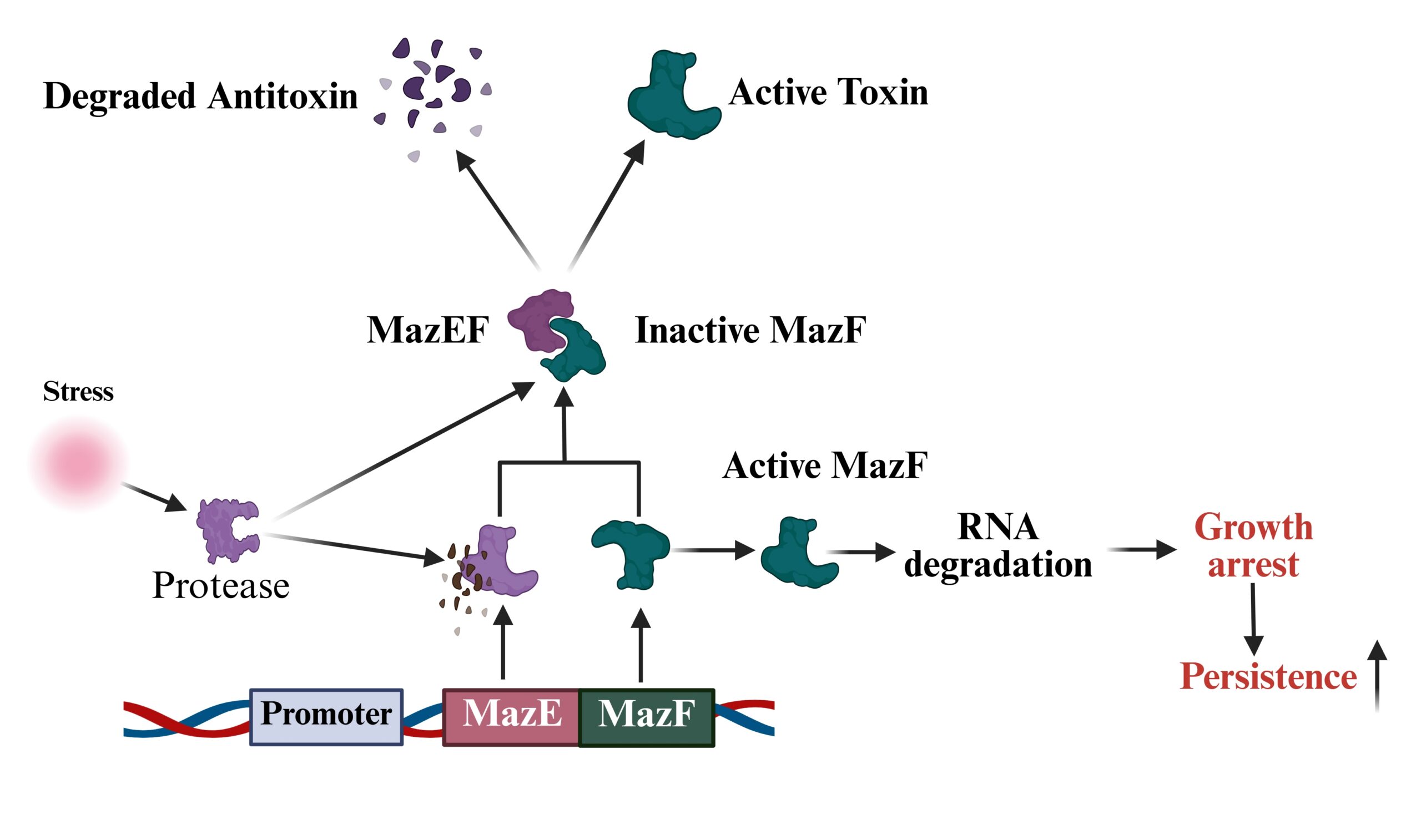
2020MazEF-rifampicin interaction suggests a mechanism for rifampicin induced inhibition of persisters Alexander C, Guru A, Pradhan P, Mallick S, et al. BMC Molecular and Cell Biology, 21(1), 73 (2020) DOI: 10.1186/s12860-020-00316-8
Analysis of Indian SARS-CoV-2 genomes reveals prevalence of D614G mutation in spike protein predicting an increase in interaction with TMPRSS2 and virus infectivity Raghav S, Ghosh A, Turuk J, Kumar S, et al. Frontiers in Microbiology, 11, 594928 (2020) DOI: 10.3389/fmicb.2020.594928
SARS-CoV-2 genome analysis of Indian isolates and molecular modelling of D614G mutated spike protein with TMPRSS2 Raghav S, Ghosh A, Turuk J, Kumar S, et al. BioRxiv (Preprint) (2020), DOI: 10.1101/2020.07.23.217430
Identification of drugs targeting multiple viral and human proteins using computational analysis for repurposing against COVID-19 Kumar S, Kumari P, Agnihotri G, Vijay A, et al. Preprint (2020), https://doi.org/10.26434/chemrxiv.12366938.v1
2019Bacterial Min proteins beyond the cell division Taviti AC, Beuria TK, Critical Reviews in Microbiology, 45(1), 22–32 (2019) DOI: 10.1080/1040841X.2018.1538932
2017MinD directly interacting with FtsZ at the H10 helix suggests a model for robust activation of MinC to destabilize FtsZ polymers Taviti AC, Beuria TK, Biochemical Journal, 474(18), 3189–3205 (2017), DOI: 10.1042/BCJ20170357
Green synthesis of multi-metallic nanocubes Patra N, Taviti AC, Sahoo A, Pal A, et al. RSC Advances, 7(56), 35111–35118 (2017) https://doi.org/10.1039/C7RA05493A
Y-shaped morphology in E. coli may be linked to peptidoglycan synthesis pathway Mallick S, Beuria TK, Canadian Journal of Biotechnology, 1(Special), 160 (2017)
2015Doxorubicin inhibits E. coli division by interacting at a novel site in FtsZ Panda P, Taviti AC, Satpati S, Kar MM, Dixit A, Beuria TK, Biochemical Journal, 471(3), 335–346 (2015)
Synthesis, characterisation and antibacterial activity of [(p-cym)RuX(L)]+/2+ complexes Tripathy SK, Taviti AC, Dehury N, Sahoo A, Pal S, Beuria TK, et al. Dalton Transactions (2015) https://doi.org/10.1039/C4DT03647F
2013In vitro evaluation of anti-infective activity of a Lactobacillus plantarum strain against Salmonella enterica serovar Enteritidis Das JK, Mishra D, Ray P, Tripathy P, et al. Gut Pathogens, 5(1), 11 (2013), DOI: 10.1186/1757-4749-5-11
2010Molecular mechanism by which the nucleoid occlusion factor, SlmA, keeps cytokinesis in check Tonthat NK, Arold ST, Pickering BF, Van Dyke MW, Liang S, Lu Y, Beuria TK, Margolin W, Schumacher MA The EMBO Journal, 30(1), 154 (2010), DOI: 10.1038/emboj.2010.285
Bacterial cytokinesis: FzlA frizzes FtsZ filaments for fission force Beuria TK, Margolin W. Current Biology, 20(23), R1024–R1027 (2010), DOI: 10.1016/j.cub.2010.10.024
2009Promoting assembly and bundling of FtsZ as a strategy to inhibit bacterial cell division: a new approach for developing novel antibacterial drugs Beuria TK, Singh P, Surolia A, Panda D, Biochemical Journal, 423(1), 61–69 (2009), DOI: 10.1042/BJ20090817
Adenine nucleotide-dependent regulation of assembly of bacterial tubulin-like FtsZ by a hypermorph of bacterial actin-like FtsA Beuria TK, Mullapudi S, Mileykovskaya E, et al. Journal of Biological Chemistry, 284(21), 14079–14086 (2009), DOI: 10.1074/jbc.M808872200
2007Totarol inhibits bacterial cytokinesis by perturbing the assembly dynamics of FtsZ Jaiswal R, Beuria TK, Mohan R, Mahajan SK, Panda D. Biochemistry, 46(14), 4211–4220 (2007) https://doi.org/10.1021/bi602573e
Interaction between cell division proteins FtsE and FtsZ Corbin BD, Wang Y, Beuria TK, Margolin W, Journal of Bacteriology, 189(8), 3026–3035 (2007)
2006Effects of pH and ionic strength on the assembly and bundling of FtsZ protofilaments: a possible role of electrostatic interactions in the bundling of protofilaments Beuria TK, Shah JH, Santra MK, Kumar V, et al. International Journal of Biological Macromolecules, 40(1), 30–39 (2006), DOI: 10.1016/j.ijbiomac.2006.05.006
2005Sanguinarine blocks cytokinesis in bacteria by inhibiting FtsZ assembly and bundling Beuria TK, Santra MK, Panda D. Biochemistry, 44(50), 16584–16593 (2005), https://doi.org/10.1021/bi050767+
A natural osmolyte trimethylamine N-oxide promotes assembly and bundling of the bacterial cell division protein, FtsZ and counteracts the denaturing effects of urea Mukherjee A, Santra MK, Beuria TK, Panda D. The FEBS Journal, 272(11), 2760–2772 (2005) DOI: 10.1111/j.1742-4658.2005.04696.x
2004Ruthenium red-induced bundling of bacterial cell division protein, FtsZ Santra MK, Beuria TK, Banerjee A, Panda D. Journal of Biological Chemistry, 279(25), 25959–25965 (2004) https://doi.org/10.1074/jbc.m312473200
2003Glutamate-induced assembly of bacterial cell division protein FtsZ Beuria TK, Krishnakumar SS, Sahar S, Singh N, Gupta K, Meshram M, Panda D. Journal of Biological Chemistry, 278(6), 3735–3741 (2003), DOI: 10.1074/jbc.M205698200 |
Group
| Details | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Current Members
Our Graduates
Lab Alumni
|
Grants
| Details | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
Extramural Grants:
|
Contacts
| Address | Fax | Office | |
|---|---|---|---|
| tkbeuria@ils.res.in | BRIC-Institute of Life Sciences, Nalco Square, Bhubaneswar-751023, India | 0091 674 2300728 | 0091 674 2302640 |
Highlights
| Details | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
Publications
Lab Group 2026
|
Positions
| Details |
|---|
PhD Opportunities: I am always looking for motivated and intellectually curious candidates with a valid CSIR–UGC JRF or equivalent national fellowship to join my research group. Candidates with a strong interest in pursuing rigorous, hypothesis-driven research are encouraged to get in touch. Postdoctoral Fellow / Research Associate / N-PDF: I welcome applications from candidates who have recently submitted their PhD thesis and are eager to engage in advanced research. If you are driven, independent, and interested in contributing to ongoing projects in my lab, please reach out with your updated CV. Enthusiastic candidates with a strong research mindset are particularly encouraged to contact me. |